Beyond Clonal Dominance: Identifying Developable Antibodies from Single-Cell In Vivo Repertoires
An end-to-end workflow for affinity-matured, liability-filtered antibody lead selection
Overview
The ultimate goal of any terminal single-cell in vivo campaign is to efficiently identify a diversified panel of developable lead candidates. However, when sequencing data is generated from pooled, sacrificed donors without longitudinal timepoints or distinct tissue compartmentalization, isolating genuine, antigen-driven clonal expansion from background noise becomes a significant computational hurdle. Platforma provides a unified, end-to-end workflow that natively supports major single-cell sequencing technologies to systematically isolate high-confidence binders.
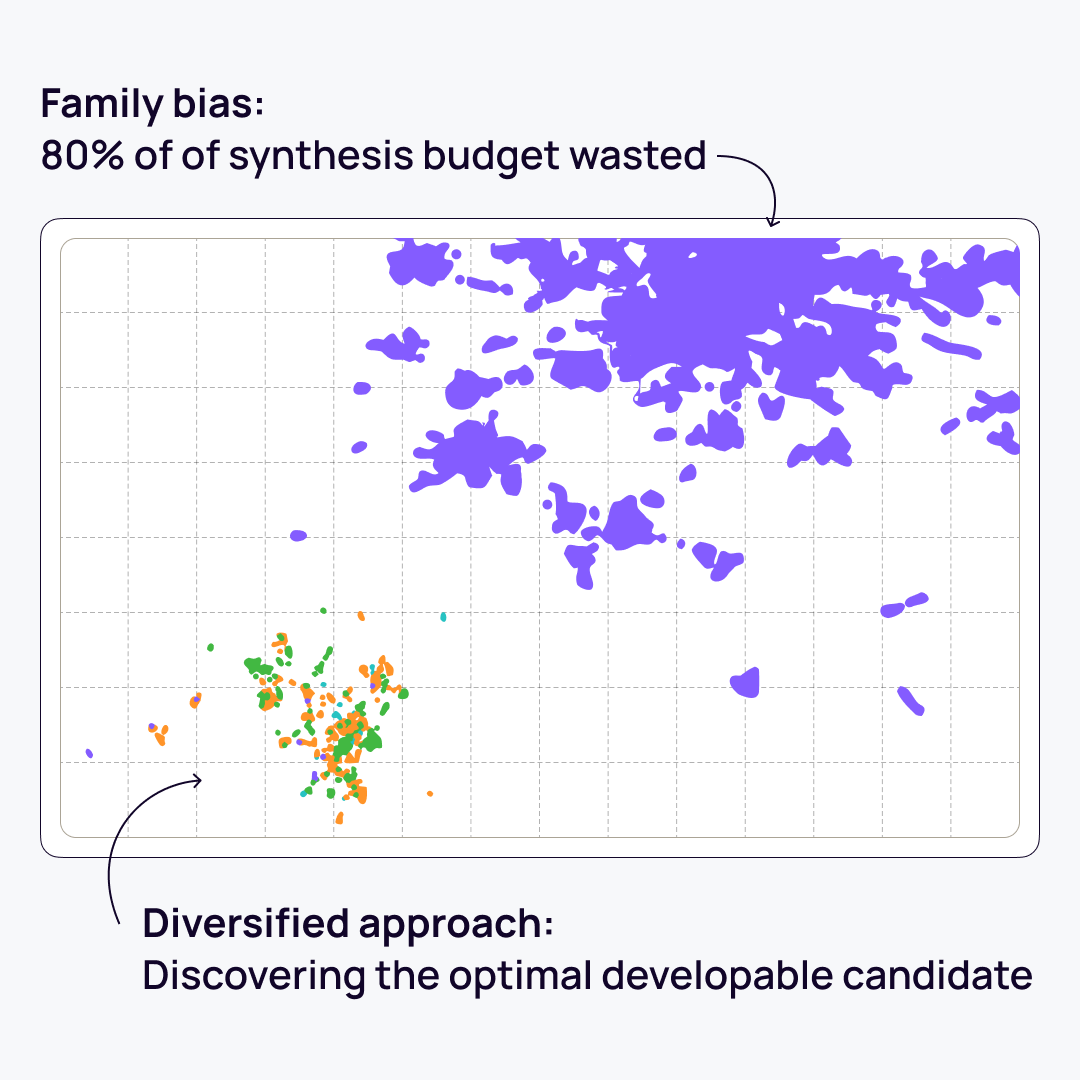
The challenge of clonal dominance
In terminal in vivo campaigns, relying purely on clone frequency is a common pitfall.
Frequency-based ranking often biases selection toward a single highly abundant lineage or non-specific expanding cells, effectively allowing one clonal family to hijack the downstream synthesis and testing budget. Furthermore, failing to enforce structural diversity across the selected cohort drastically increases the risk of developmental failure due to shared physicochemical liabilities.
Methodology
Platforma shifts the paradigm from raw frequency ranking to a multi-dimensional evaluation. By pairing precise MiXCR clonotyping with advanced paratope grouping, liability screening, and somatic hypermutation (SHM) metrics, it algorithmically balances clonal abundance with structural maturation. This enables teams to mine complex repertoires and prioritize variants with optimal immunological fitness.
The workflow integrates four key pillars:
Clonotyping: Leverages MiXCR for exact V(D)J assembly, error correction, and extraction of critical SHM maturation profiles.
Dual-Tier Clustering: Enforces structural diversity by grouping variants via sequence similarity and predicted binding interfaces.
Liability Screening: Scans the repertoire for physicochemical weaknesses to filter out severe aggregation/cleavage motifs prior to synthesis.
Algorithmic Scoring: Employs In Vivo Score to rank candidates based on a composite of clonal expansion and affinity maturation.
Building a Diversified Lead Panel
Platforma’s reproducible decision matrix allows teams to execute either an unbiased global panel diversification or targeted seed-based NGS mining within the same dataset.
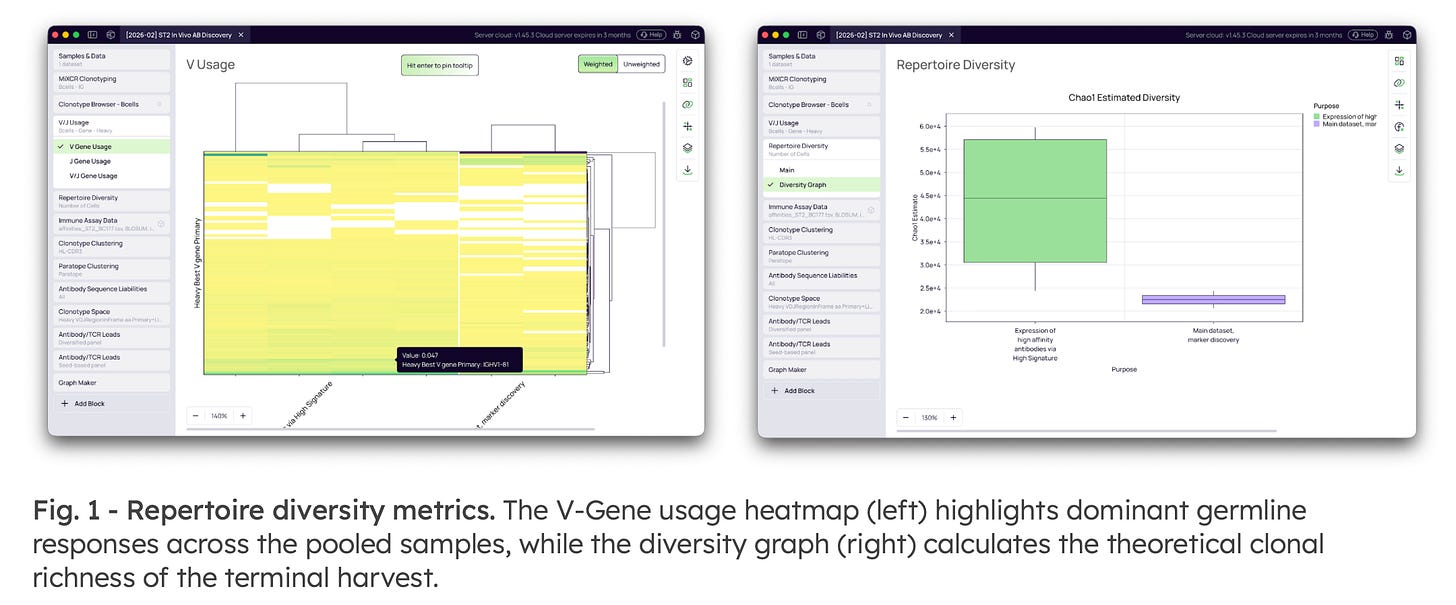
1. High-Precision Annotation & Repertoire Characterization
The workflow utilizes the MiXCR engine to perform exact V(D)J assembly and extract full-length sequences. It automatically calculates comprehensive SHM metrics—such as CDR mutation fractions—which are critical for identifying affinity-matured clones. Researchers can also assess macro-level biological responses by calculating theoretical clonal richness (Chao1 Estimate) and visualizing dominant germline scaffolds.
2. Lineage And Paratope Clustering
To enforce structural diversity, unique sequence variants must be grouped into biological families. The pipeline performs sequence-based clustering to group functional somatic variants (Lineage Grouping), followed by predictive paratope clustering. This ensures that selected sequences represent distinct functional binding interfaces rather than redundant somatic cousins.
3. Sequence Liability Screening
To protect the downstream synthesis budget, the algorithm scans the repertoire for physicochemical weaknesses that compromise developability. Candidates are evaluated for risks such as fragmentation, isomerization, N-linked glycosylation, and deamidation, allowing teams to filter out “High Risk” severe aggregation or cleavage motifs before lead selection.
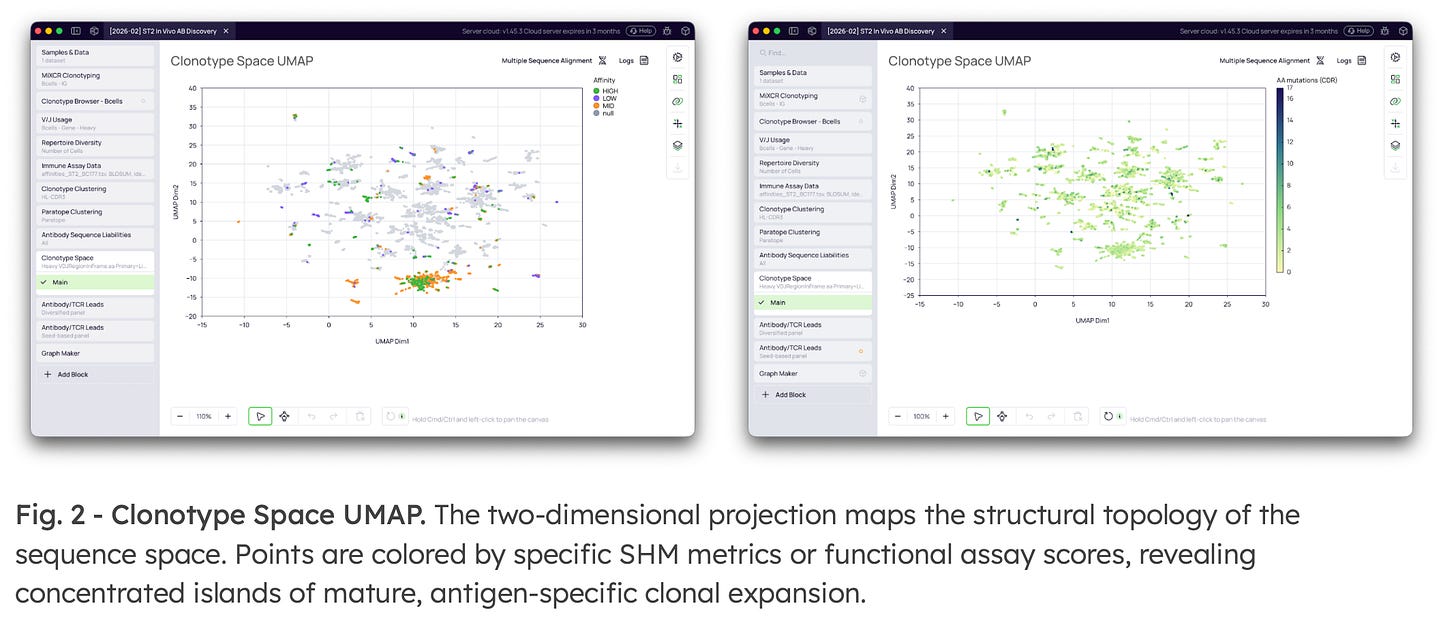
4. Assay Data Integration & UMAP Visualization
Platforma bridges computational discovery with wet-lab validation. Users can import functional hit datasets (e.g., K.D., ELISA scores) and use fuzzy matching to capture naturally occurring somatic variants of baseline seeds. The entire multidimensional repertoire is then projected onto a UMAP landscape.
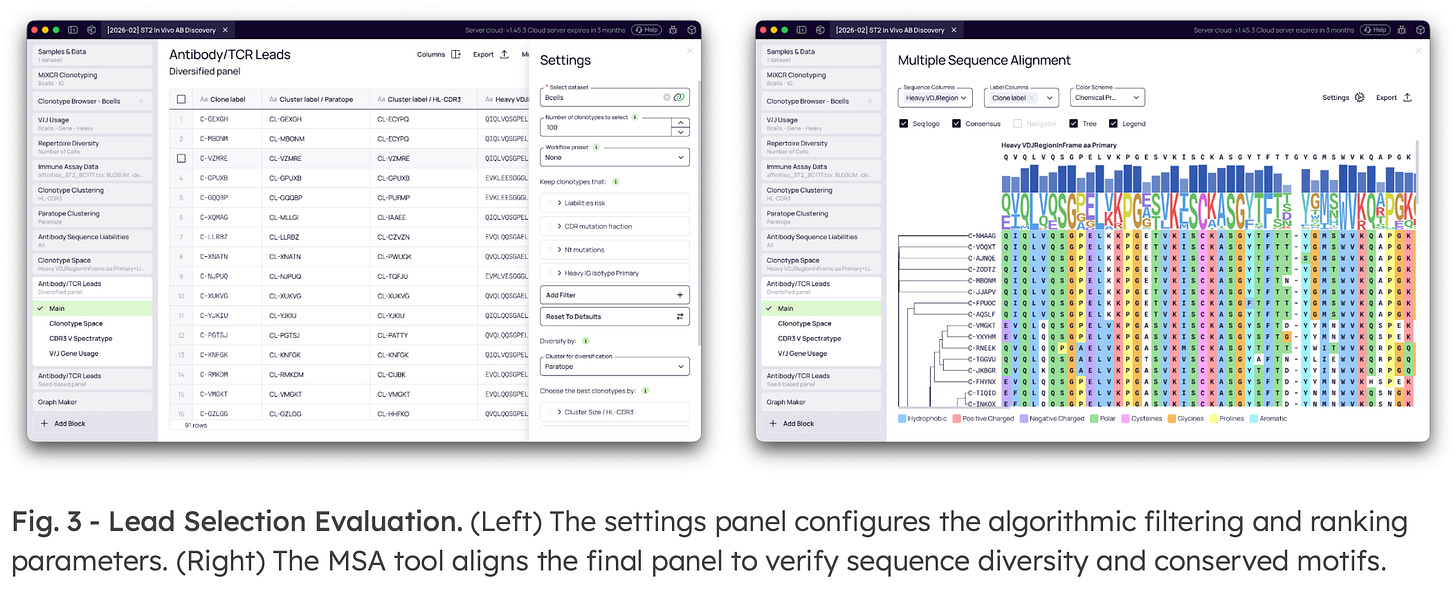
5. Lead Selection Fitting
The core of Platforma’s lead selection is the In Vivo Score—a multi-parametric algorithm that substitutes raw frequency ranking with balanced sequence selection. The score mathematically evaluates to: 40% (clonal frequency) + 35% (CDR mutation fraction) + 25% (nucleotide mutations).
Using this funnel, teams can isolate class-switched cells, reject unmutated background clones, require candidates to belong to an expanded clonal family, and force the sampling of distinct paratope clusters to generate a globally diversified, developable panel.
Conclusion
Platforma’s computational workflow transforms raw, endpoint single-cell in vivo data into a highly specific, developable panel of affinity-matured variants. By systematically evaluating baseline abundance alongside targeted somatic hypermutation rates and physicochemical liabilities, discovery teams can avoid the pitfalls of frequency bias, maximize topological coverage, and significantly reduce downstream developmental risk.