Beyond Fold Change: Enrichment Quality, Binding Specificity, and Cross-Antigen Analysis
Computationally identify antigen-specific leads without additional wet-lab experiments
Introduction
In display-based discovery campaigns, finding clones with a high fold change across selection rounds is necessary but not sufficient. A clone might expand simply because it binds a carrier protein, the solid support, or grows efficiently in the host organism—not because it binds the target antigen. Historically, separating true antigen-specific binders from non-specific expanders has required costly and time-consuming downstream wet-lab validation.
The Limits of Fold Change
Relying solely on the traditional criterion—ranking clusters by the highest fold change—conflates three biologically distinct phenomena:
genuine antigen-specific selection
non-specific enrichment of fast-growing clones
background amplification of library-dominant sequences
While aggressive washing attempts to minimize this noise, real-world campaigns rarely achieve a perfect ideal. Without a computational means of discrimination, non-specific binders routinely occupy the top of the enrichment list.
Solution: Multidimensional Enrichment Analysis
The Clonotype Enrichment analysis in Platforma addresses this gap computationally. By easily integrating negative control pannings (such as rounds against BSA or a surface without antigen) , the platform evaluates candidates beyond simple log₂ fold change. It classifies each antibody family along two independent axes:
Enrichment Quality: How consistently and progressively a clone was selected across rounds.
Binding Specificity: Whether enrichment was driven by the target antigen or by non-specific binding to controls.
Additionally, its modular architecture enables cross-antigen specificity analysis: by running parallel enrichment blocks for different targets, teams can identify monospecific binders or cross-reactive candidates from a single campaign without additional wet-lab experiments. Operating on cluster-level data, these classifications feed directly into automated candidate delivery.
Advanced Capabilities of the Enrichment Analysis
This workflow details a computational workflow for distinguishing true binders from background noise by evaluating antibody families based on cluster-level abundance, enrichment quality, and binding specificity.
1. Configuring enrichment on antibody clusters
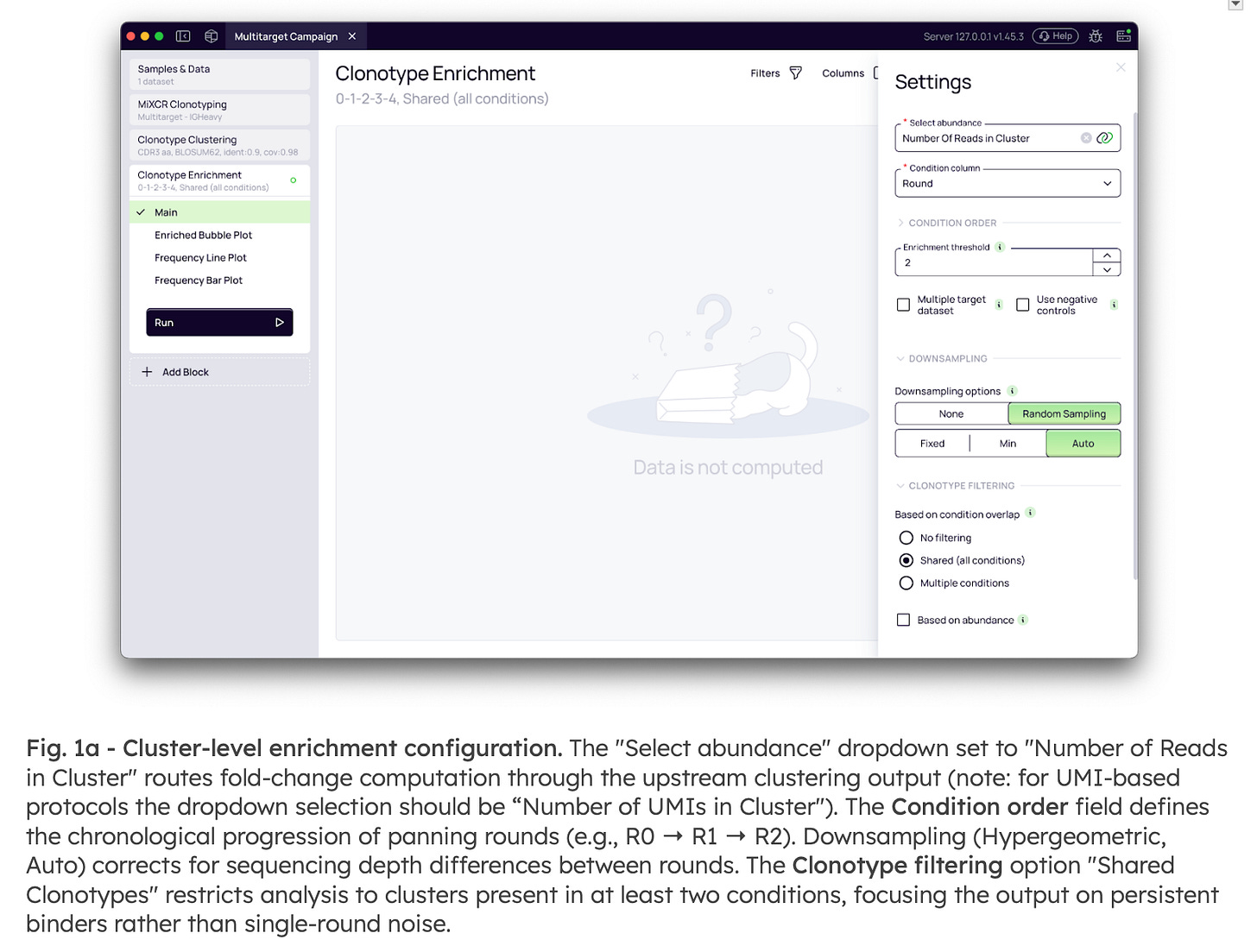
Analysis begins by linking directly to the cluster-level abundance generated upstream. By aggregating the signal across sequence variants within a single lineage, the platform recovers rare binders that might otherwise fall below the detection threshold if analyzed individually. The workflow automatically accounts for the chronological progression of panning rounds, normalizes for sequencing depth differences via downsampling, and filters for persistent binders to eliminate single-round noise (Fig. 1a)
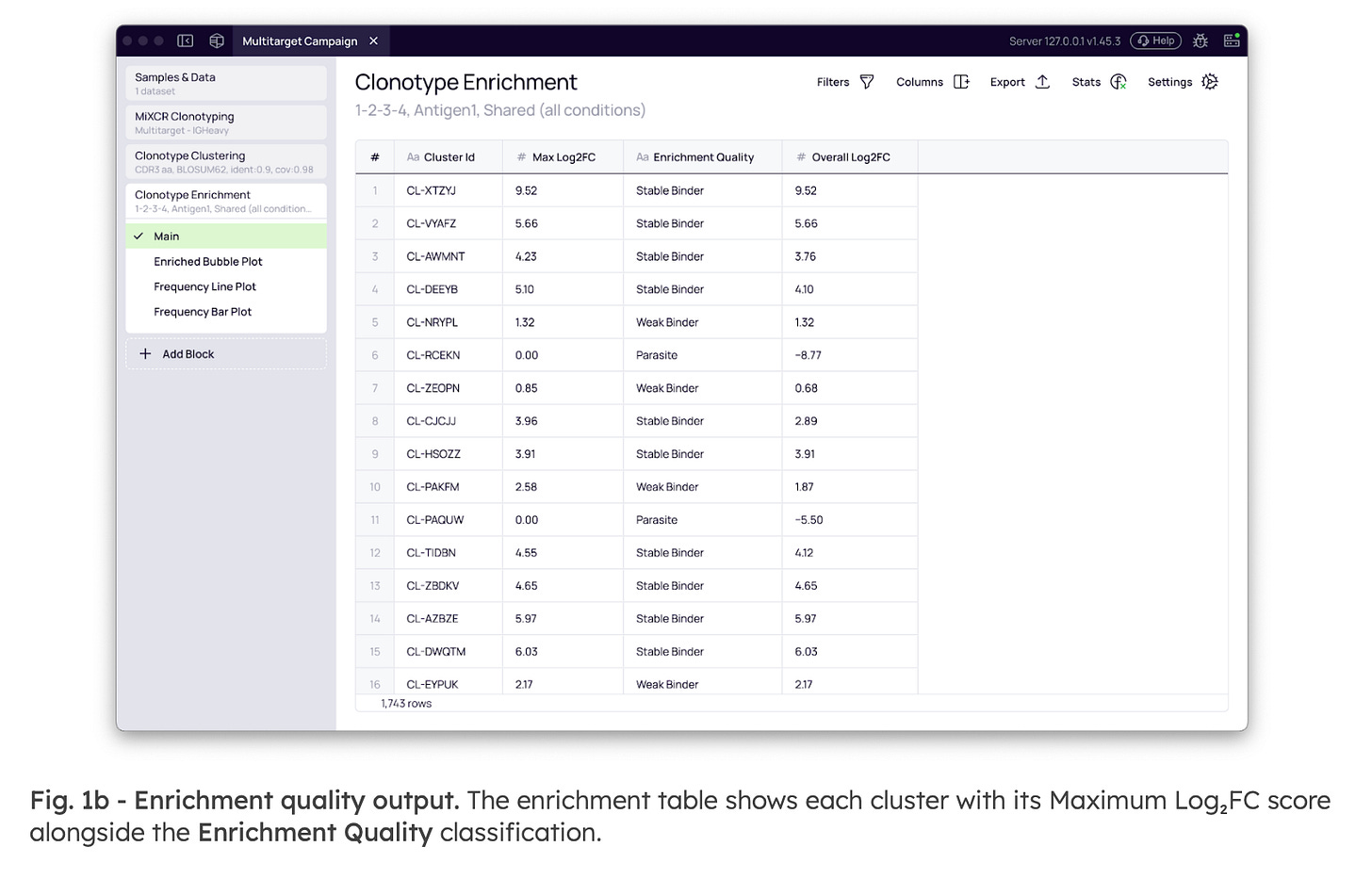
To provide immediate interpretive context beyond a raw Log2 Fold Change score, the output classifies each cluster’s trajectory (Fig. 1b). This allows teams to distinguish candidates with a reliable monotonic response from those with ambiguous trajectories before committing to synthesis. The Enrichment Quality labels include:
Stable Binder: Consistently enriched in every round.
Rescuer: Enriched in later rounds after an initial dip.
Parasite: High early but declining.
Weak Binder: Marginally enriched.
2. Binding specificity classification
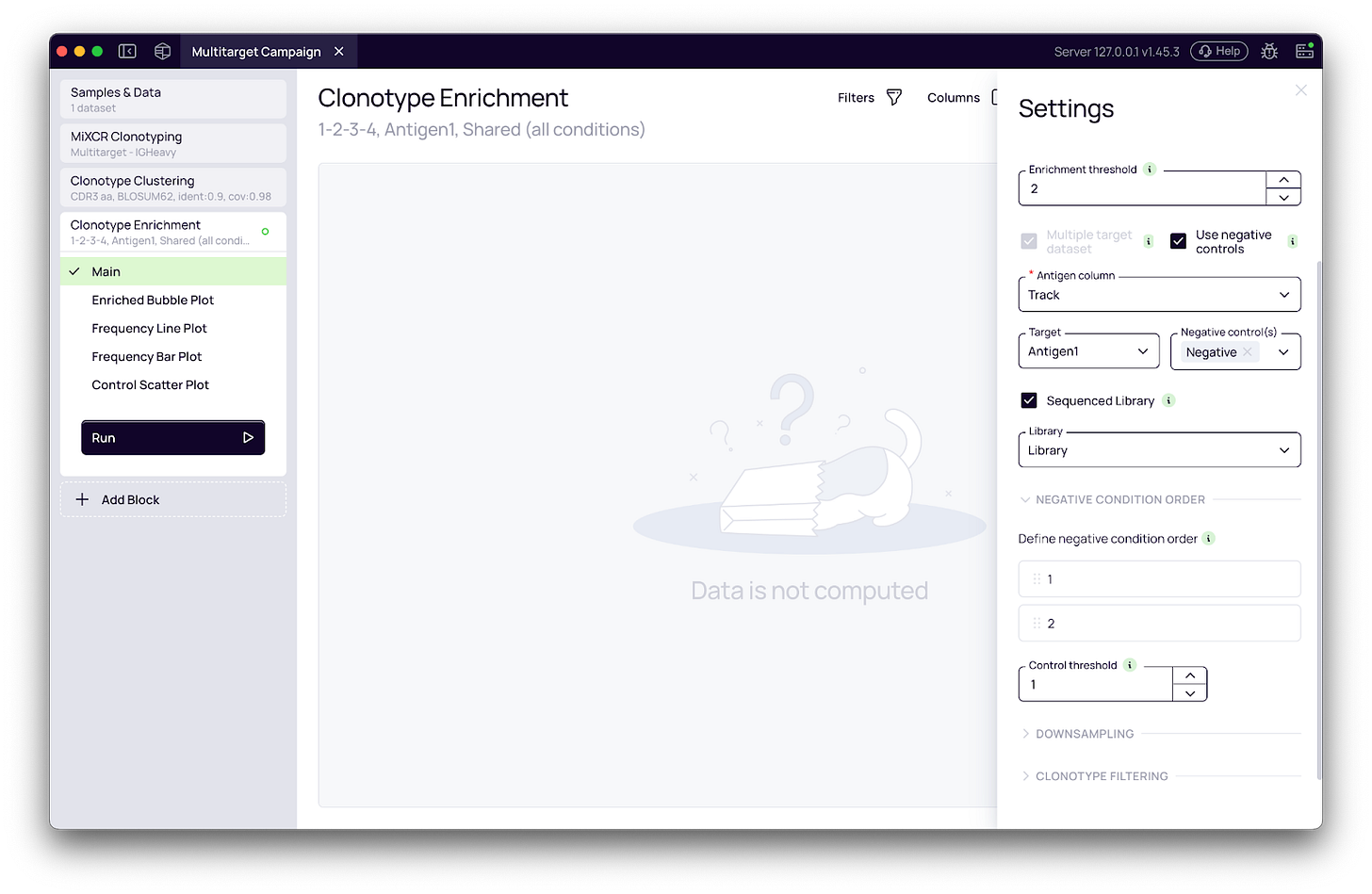
When an experiment includes negative control pannings, the workflow enables a second critical layer of analysis. By mapping sample metadata to distinct target and negative control conditions (e.g., BSA or bare plastic), the platform independently computes fold change for each environment (Fig. 2a). This is flexible enough to support campaigns where target and control pannings follow completely different round structures.
Based on where the enrichment signal originates, the workflow assigns a Binding Specificity label to every cluster:
Antigen-Specific: Enriched in target rounds, not in controls.
Non-Specific: Enriched in both target and control rounds.
Negative-Control: Enriched exclusively in controls—acting as a clear liability flag.
Not-Enriched: No significant enrichment in either condition.
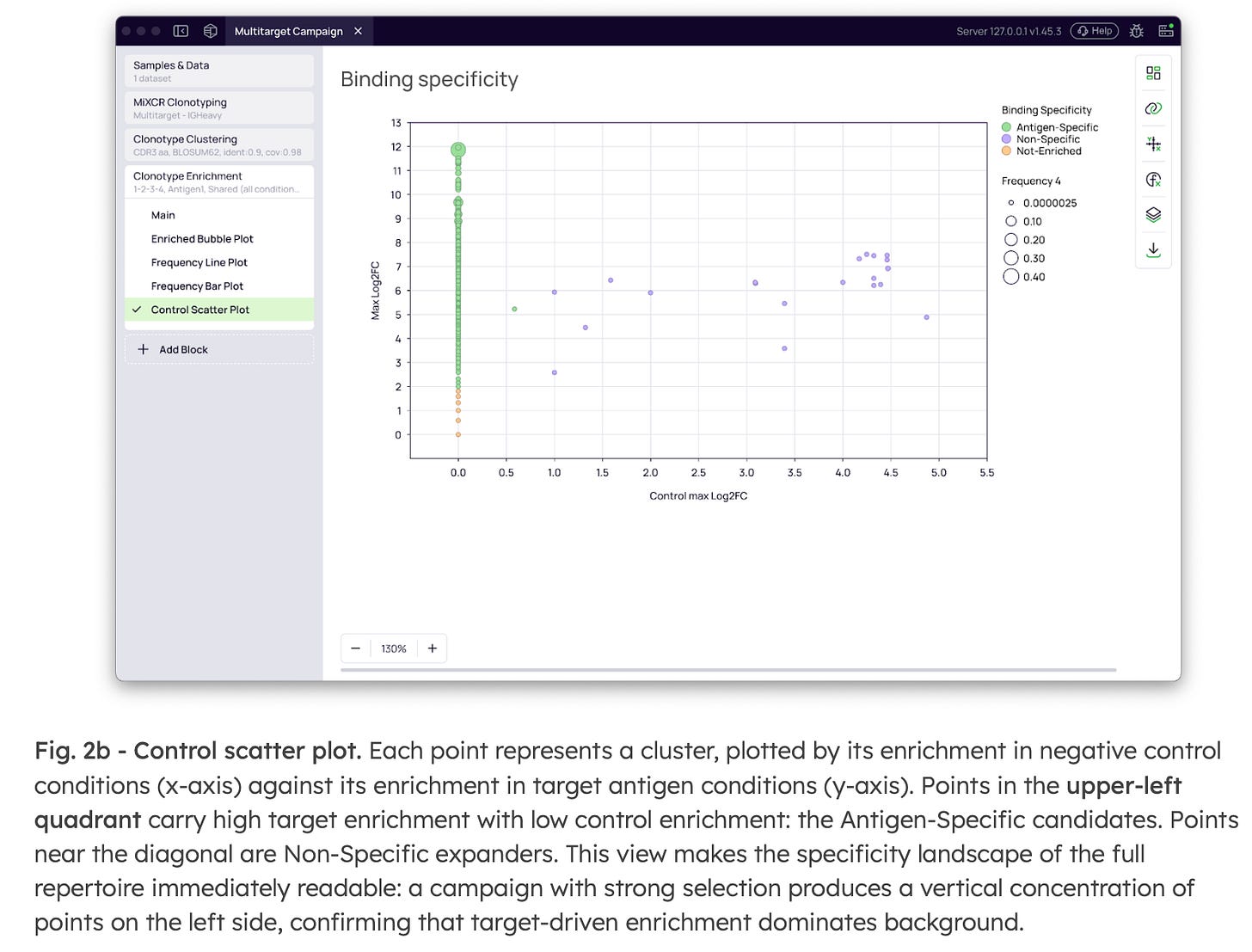
These labels are exported as structured columns, immediately available for downstream lead selection. Furthermore, this data generates a Control Scatter Plot (Fig. 2b) that makes the specificity landscape of the full repertoire instantly readable. Antigen-specific candidates isolate cleanly into the upper-left quadrant (high target enrichment, low control enrichment), while non-specific expanders track predictably along the diagonal
Fig. 2a - Antigen and negative control configuration. The “Target selection” section activates binding specificity analysis. The Antigen column dropdown selects the metadata column mapping each sample to its antigen. The Target antigen dropdown identifies which value represents the selection target; the Negative controls multi-select captures the irrelevant antigens used as specificity references (e.g., BSA, Plastic). The condition order for control rounds is specified independently of the main target condition order, supporting campaigns with different round structures for target and control pannings.
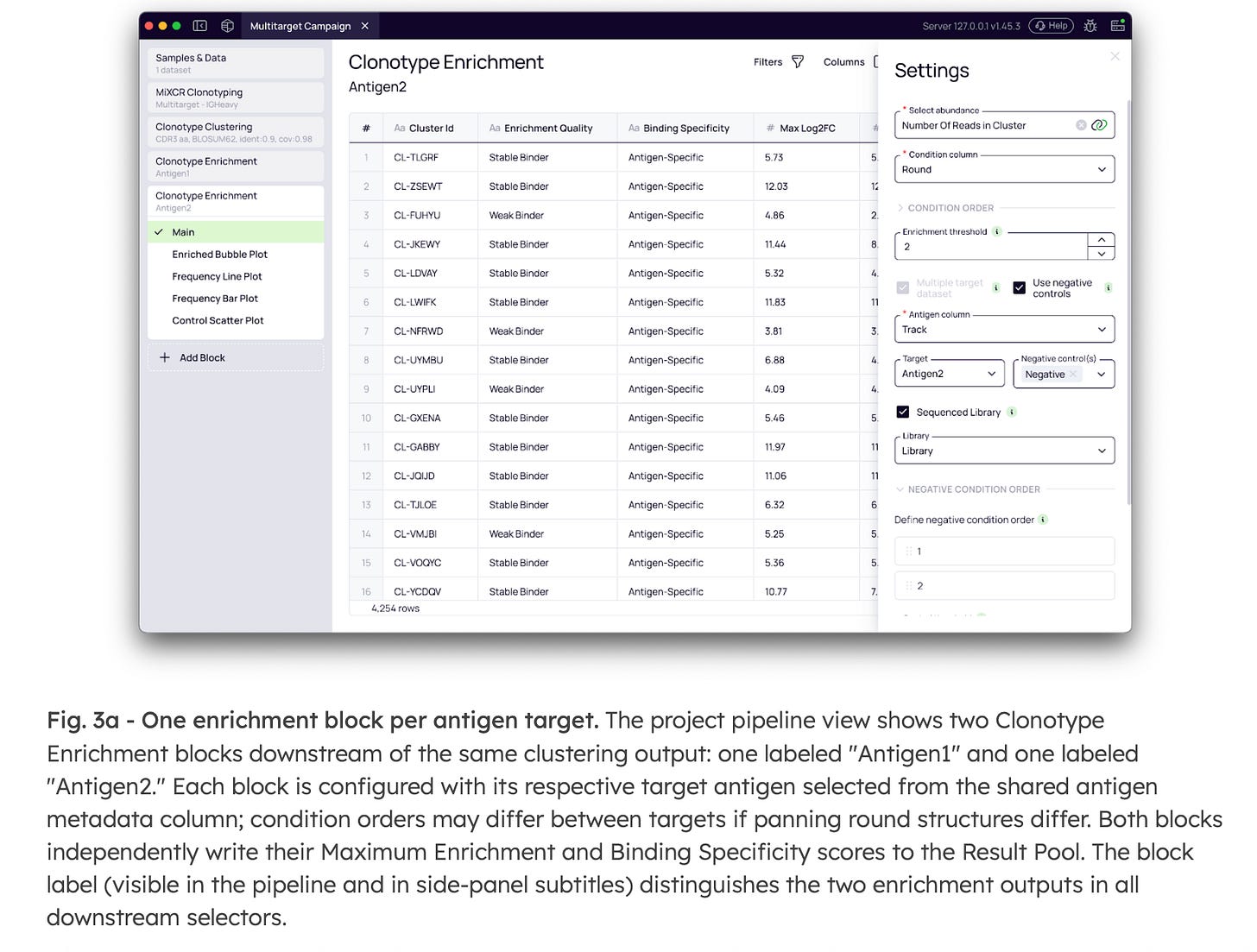
3. Cross-antigen specificity analysis
Multi-antigen campaigns—where a single library is panned against two or more distinct targets—introduce a critical question: which clusters are selective for a single antigen, and which are broadly cross-reactive? Platforma’s modular architecture resolves this computationally, requiring no additional sequencing or wet-lab experiments.
By running parallel Clonotype Enrichment analyses for each target antigen within the same project, the platform independently calculates enrichment scores and specificity classifications for each target (Fig. 3a). Because these parallel analyses feed into a unified data environment, the Lead Selection step can query multiple enrichment profiles simultaneously (Fig. 3b).
By applying specific filters within the Lead Selection, you can directly identify your exact desired binding profiles:
Monospecific Binders: Isolated by requiring a cluster to be “Antigen-Specific” for one target while remaining “Not-Enriched” or “Non-Specific” for all others.
Cross-Reactive Candidates: Identified by requiring strong, target-specific enrichment across multiple parallel analyses.
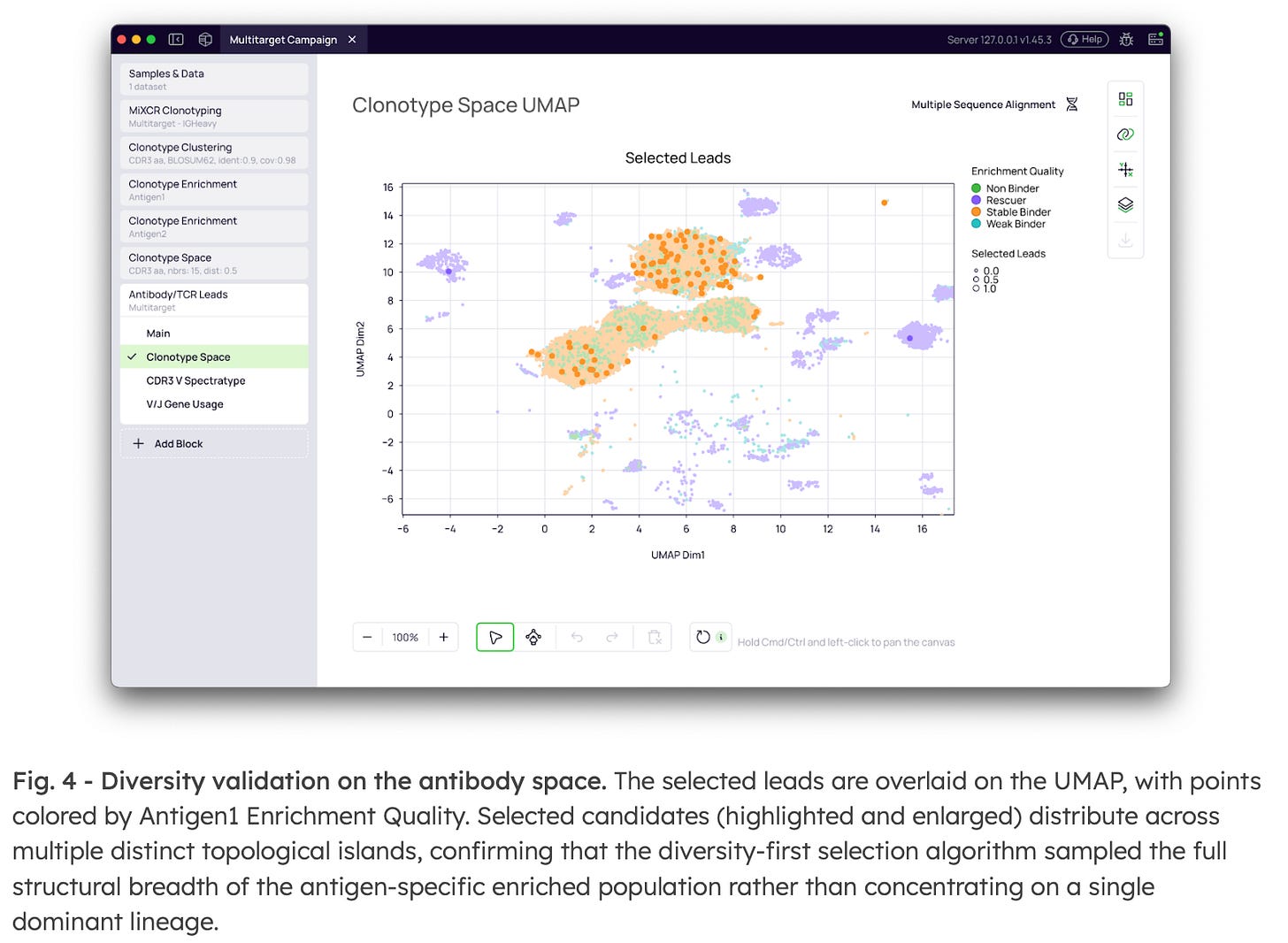
4. Spatial validation of the candidate panel
The final selected leads are overlaid on the UMAP projection of the full repertoire, colored by Binding Specificity from each enrichment block. Monospecific candidates occupy distinct topological regions from cross-reactive clusters, providing spatial confirmation that the selection criteria isolated the intended specificity profile. The combination of Enrichment Quality, Binding Specificity, and cross-antigen filter settings converts what was previously a manual, multi-step analytical process into a single, reproducible configuration within one project.
Conclusion
The advanced capabilities of the Clonotype Enrichment block transform enrichment analysis from a single-metric ranking into a multi-dimensional characterization of each antibody family’s selection behavior. By evaluating cluster-level data—which aggregates signal across sequence variants to rescue rare, low-abundance binders—discovery teams can confidently characterize:
Enrichment Quality: Distinguishing stable, progressive binders from transient or declining clones.
Binding Specificity: Separating genuine antigen-driven enrichment from non-specific background amplification using negative controls.
Cross-Antigen Profiles: Automatically identifying monospecific and cross-reactive candidates directly from NGS data without additional experiments.
These classifications flow seamlessly into automated candidate selection, serving simultaneously as stringent hard filters and ranking criteria. This workflow ensures that every lead candidate delivered for synthesis carries a quantitative, data-driven justification that is fully traceable to the original sequencing data.